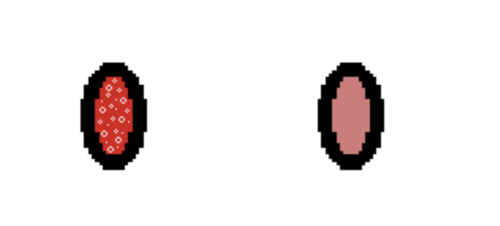Biomedical Forensics (Virtual)
Objective
The objective of this experiment is to use biomedical forensic techniques, including extracting DNA, fingerprinting, identifying foreign substances, and performing blood typing to investigate a crime scene. The DNA will be extracted using the basic biochemical techniques for isolating, purifying, and digesting DNA molecules. The drug testing will be completed by observing chemical reactions using simulated reagents. A blood typing simulation kit will be used to test the blood.
Overview
Most people learn about biomedical forensics from TV shows that, in fact, misrepresent this branch of science. It is true that biomedical forensic methods are commonly used in criminal or civil cases, but some of these techniques also have applications in medicine in the diagnosis and treatment of diseases and injuries, in product safety and analyzing how and why products and systems affect users, and other engineering fields. In criminal law, these techniques are used to identify suspects in criminal cases and to exclude individuals as suspects. DNA testing, in particular, is increasingly used to prove the innocence of people who have been wrongfully convicted of a crime.
The Structure of DNA
Deoxyribonucleic acid (DNA) is found in almost all living organisms. These organisms can be as simple as single-celled bacteria or as complex as a multi-celled human; the human body contains approximately 50 trillion cells. There are two different types of cells: prokaryotes and eukaryotes. Prokaryotic cells do not have a nuclear membrane and so do not have a distinct nucleus. An example of prokaryotic organisms are bacteria. Only eukaryotic cells, which are found in plants and animals, will be considered in this lab. Eukaryotic cells have a distinct, membrane-bound nucleus that isolates the DNA from the rest of the cell. Plant cells are different from animal cells in structure and cellular contents. Only plant cells will be used in this experiment.
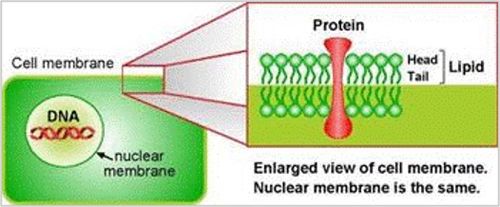
Plant cells are surrounded by a cell wall that has high mechanical strength and protects the cell. The cell wall lies directly beneath the plasma membrane (Figure 1) that separates the interior of the cell from the outside environment. Cytosol is found within the plasma membrane.
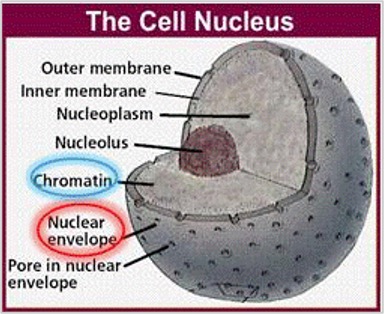
The various cell organelles that perform specialized functions for the cell, including the nucleus, are found within the cytosol. The nucleus (Figure 2) houses DNA in the form of chromatin, which is the building block for chromosomes.
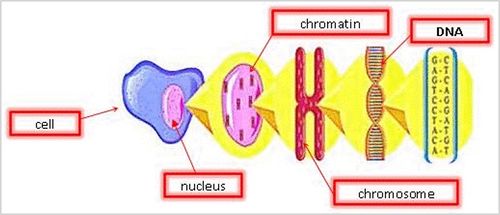
Chromatin is the active form of DNA in the cell when it is not preparing for cell division. It is comprised of DNA wrapped around protein particles called histones, which help pack and order the DNA into structural units (Figure 3).
The DNA molecule is a double-helical polymer consisting of a sugar-phosphate backbone with nitrogenous bases running perpendicular to the backbone. These bases, often represented by letters A (adenine), G (guanine), C (cytosine), and T (thymine), are the elementary components making up coded genetic information (Figure 4). The base sequence acts as the instruction manual for the cell directing it on how to make proteins and other important molecules that an organism needs to survive and function.
DNA Extraction
A process that will be used in this experiment is to extract the DNA from a fruit sample. Some knowledge of DNA extraction is needed to do this. The DNA extraction process is a fairly simple biochemical procedure that can be divided into three major steps: breaking open the cell wall, destroying membranes within the cell, and precipitating the DNA out of the solution.
The first step in DNA extraction is to break the cell wall (cell lysis) to expose the DNA within the cell. Plant cells have a very rigid external structure — the cell wall — that protects the intracellular components. It is very rigid and acts as a protector and filter for substances moving in and out of the cell. It is made of cellulose, which is the insoluble substance that makes wood hard and durable. To destroy the cell wall, a mechanical method is used to break apart the cellulose molecules. In this experiment, the fruit sample will be mashed manually.
The second step in DNA extraction is to destroy the plasma membrane within the cell. The cell's plasma membrane is made of phospholipid bilayers that are made of fat. To disrupt them, the mesh of fat molecules is broken up with a surfactant (soap). The structure of soap is very similar to the structure of fat and grease and lowers the surface tension between two fluids when acting as a surfactant.
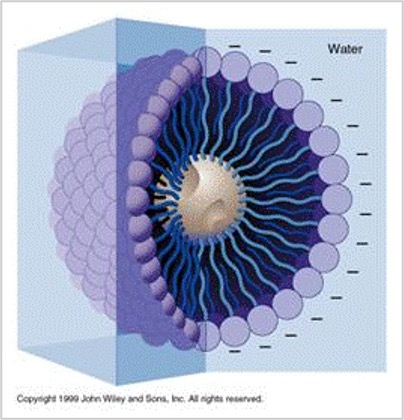
A soap molecule has two parts: a head and a tail. The head is polar and is attracted to water (hydrophilic) while the tail is non-polar and attracted to oil and fat (hydrophobic). When soap molecules are in water, they group themselves into micelles that have a roughly spherical structure in which all the polar heads point outwards (toward water) and all the non-polar tails point inwards at the center of the sphere (away from the water) (Figure 5).
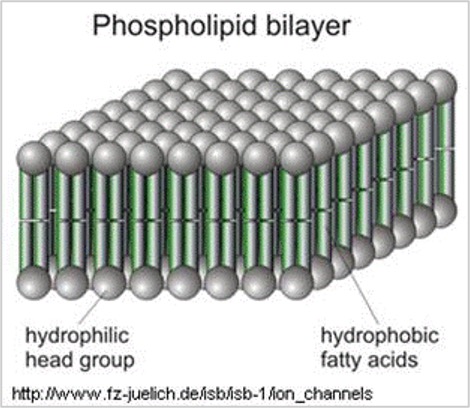
The soap molecules effectively trap the fat molecule inside the micelle and dissolve the cell membranes. The molecules in the phospholipid bilayer (Figure 6) also contain molecules that are made up of a hydrophilic head and a hydrophobic tail. The soap molecules orient themselves so that their head associates with the tail of the phospholipid bilayer. In this way, the soap breaks up the bilayer molecule by molecule.
The last step in DNA extraction is precipitating the DNA. When the plasma membrane is successfully disrupted, the DNA is released from the cells into the solution along with protein molecules and other cellular miscellany (Figure 6). With the cell's contents mixed into a solution, the DNA can be separated from the rest of the contents. This process is called precipitation. Salt is used because it disrupts the structure of the proteins and carbohydrates found in the solution. Also, the salt provides a favorable environment to extract the DNA by contributing positively-charged sodium ions that neutralize the negative charge of DNA.
After the addition of salt and soap, the extracted DNA cannot be seen as it is too small to distinguish from the rest of the solution. To aid in making the DNA visible, alcohol is added since it cannot dissolve the DNA. A translucent white substance will begin to form at the top: this is the DNA (Figure 7). Once the precipitate is thick enough, it can be spooled out. This simple procedure is a rough extraction process that needs further purification before it can be successfully run on a gel for analysis.
Fingerprints
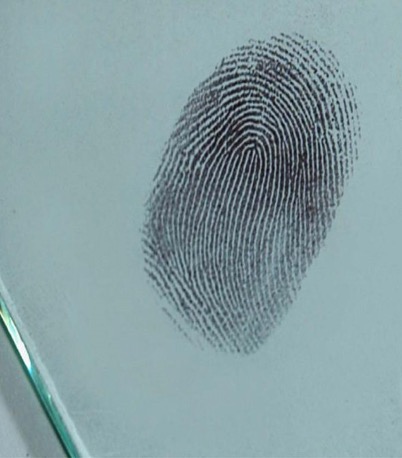
Fingerprints are unique patterns consisting of friction ridges (raised) and furrows (recessed) that appear on the pads of the fingers and thumbs, They are often used for identification purposes in forensics. Prints from palms, toes, and feet are also unique, but these are used less often for identification so this guide focuses on prints from the fingers and thumbs.
The fingerprint pattern, such as the print left when an inked finger is pressed onto paper, is made from the friction ridges on that finger. Friction ridge patterns are grouped into three distinct types — loops, whorls, and arches — each with unique variations depending on the shape and relationship of the ridges.
Loops are prints that recurve back on themselves to form a loop shape (Figure 8). Divided into radial loops (pointing toward the radius bone, or thumb) and ulnar loops (pointing toward the ulna bone, or pinky), loops account for approximately 60% of pattern types.
Whorls form circular or spiral patterns, like tiny whirlpools (Figure 9). There are four groups of whorls: plain (concentric circles), central pocket loop (a loop with a whorl at the end), double loop (two loops that create an S-like pattern), and accidental loop (irregularly shaped). Whorls make up about 35% of pattern types.
Arches create a wave-like pattern and include plain arches and tented arches (Figure 10). Tented arches rise to a sharper point than plain arches. Arches make up about 5% of all pattern types.
Blood Typing
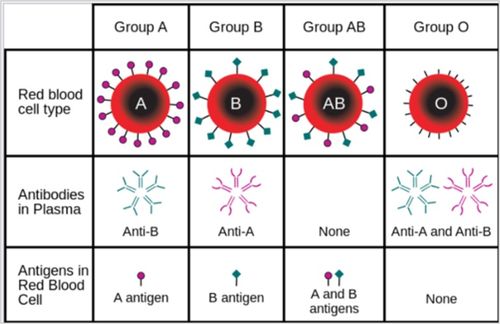
Even though all red blood cells are made of similar elements, not all blood cells are alike. There are eight different types of blood, each based on the presence or absence of three types of antigens (A, B, and Rh) (Figure 11). Antigens in general induce an immune response to foreign substances in the body. The A and B antigens determine the type of blood while the Rh antigen determines if the blood is positive or negative with the Rhesus factor, which is associated with hemolytic diseases and incompatible blood transfusions.
- The ABO Blood Group System
- Group A has only the A antigen on red cells
- Group B has only the B antigen on red cells
- Group AB has both A and B antigens on red cells
- Group O has neither A nor B antigens on red cells
Positive (+) blood types have the Rh factor and negative (-) blood types do not have the Rh factor.
Materials and Equipment
- Computer with internet access
- Paper
- Pencil with graphite
- Clear packaging tape
Procedure
Forensic Academy
Basic training in three forensic techniques will be completed before the crime scene investigation is conducted.
1. DNA Extraction
The procedure below was previously performed and videoed by TAs. Watch DNA Extraction Part 1 and DNA Extraction Part 2. Download the video if it does not play in the browser.The steps are outlined for your information only.
- Obtain distilled water in a beaker.
- Add 2 tsp (10 ml) of dish soap to the water.
- Stir in ¼ tsp salt and mix until the salt dissolves. This is the extraction mixture.
- Place one strawberry into a Ziploc bag.
- Pour the extraction mixture into the bag with the strawberry.
- Remove as much air from the bag as possible and seal it closed.
- Mash the strawberry inside the bag manually. Crush any large pieces.
- Pour the resulting strawberry pulp and extraction mixture through a strainer and into a beaker.
- Use a plastic spoon to press the mashed bits of strawberry against the strainer forcing as much of the mixture as possible into the container.
- Pour the extraction mixture into a test tube. This will help isolate the DNA on the surface of the mixture.
- Add 1 tsp (5 ml) of the chilled isopropyl alcohol to the solution (request the isopropyl alcohol from a TA) and hold the mixture at eye level. Look for the separation of DNA that shows up as a white layer on the surface of the mixture.
- If the separation of DNA cannot immediately be seen, set the small container aside for a few minutes and examine it again later.
2. Fingerprinting
Fingerprinting will be performed by using at-home materials.
- Draw a filled-in rectangle (approximately 1” x 2") on paper using a graphite pencil. Fill in the square completely and as dark as possible to ensure a clear fingerprint. Alternatively, a pen or marker can be used if a pencil is unavailable.
- Press a thumb on the rectangle and hold for approximately 5 seconds.
- To obtain the print, place the thumb on a piece of clear packaging tape with a bit of force and then lift it. Place the tape on a white sheet of paper. Alternatively, if packaging tape is unavailable, the thumb can be applied directly to the sheet of paper.
- Closely observe the fingerprint and identify the fingerprint type. Refer to the Fingerprints section in the Overview to identify the type of fingerprint.
- Take a picture of the fingerprint as your Lab Notes.
3. Drug Testing
Drug testing will be performed with a virtual simulation.
- Open the virtual drug testing simulation here.
- Note the microcentrifuge tubes labeled A, B, C, D, and E that contain the drug samples.
- Note the five drug reagents in the beakers. The reagents test for the following drugs.
- Drug A: Cannabis
- Drug B: Heroin
- Drug C: MDMA (Ecstasy)
- Drug D: Cocaine
- Drug E: Lysergic Acid Diethylamide (LSD)
- Place a droplet of the cannabis reagent in the A-labelled drug sample by clicking and dragging the pipette. If droplets of the wrong drug sample or reagent are collected, simply click and drag the pipette to the waste beaker. Remember to clean the pipette before and after using each drug reagent. Simply click and drag the pipette to the water beaker and similarly empty the water from the pipette in the waste beaker.
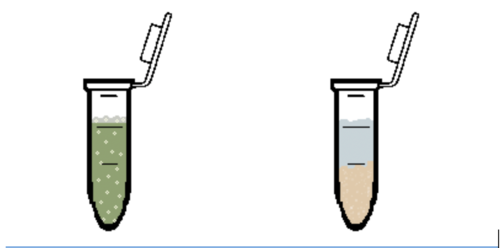
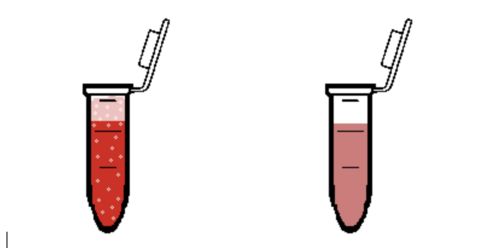
- Observe the microcentrifuge tube. If a reaction is observed from adding the reagent, the tube contains that drug (Figure 12). Record which reagents react in your Lab Notes.
- Repeat Steps 4 and 5 for each of the drug reagents.
Congratulations, the Forensic Academy is now completed and the crime scene investigation can begin.
Crime Scene Investigation
It was a warm and stormy night on July 23, 2017. The EG1004 office was quiet as one TA was pulling a late night shift to prepare the classroom for the next morning’s lab. There was a loud crash in the background. The room went dark. The next morning, there was police tape barring entry to the entire room. Someone had murdered Ruhit Roy, and now his killer must be found. At the crime scene, police discovered a pool of blood, along with the suspected murder weapon — a shard of glass — near the body. They have narrowed down the suspects to the 10 people who had access to the office that late at night.
Download the Suspect Information Sheet to get started on the investigation.
1. Drug Testing
To collect important information on the murder, the victim’s blood will be tested to determine the drug that killed him.
- Open the virtual drug testing simulation here. Go to the Crime Scene by clicking the setting at the bottom right of the page.
- Click and drag the pipette to put a sample of Roy’s blood into each of the five microcentrifuge tubes.
- Following the procedure used in the Forensics Academy, use the drug reagents to determine which drug was injected into Roy’s bloodstream by the killer (Figure 13). Remember to clean the pipette before and after using each drug reagent. Simply click and drag the pipette to the water beaker and similarly empty the water from the pipette in the waste beaker.
- Record the drug that was used on the victim in your Lab Notes.
2. Blood Typing
- Open the virtual simulation here to obtain the blood samples of all 10 suspects and of the victim along with their respective A, B, and Rh serum capsules.
- Note which suspect is being tested at the top of the page. Click and drag the pipette to administer Suspect 1’s blood into all three ovals of the blood test tray. Click and drag the pipette to the waste beaker to empty it.
- Using the pipette, administer one drop of the A serum into the oval labeled A, one drop of the B serum into the oval labeled B, and one drop of the Rh serum into the oval labeled Rh. Remember to clean the pipette before and after using each serum and blood sample. Simply click and drag the pipette to the water beaker and similarly empty the water from the pipette in the waste beaker.
- If the blood sample exhibits a reaction to the serum, then the blood type is positive for that type. If the liquid appears to be more diluted (more watery or thin), then the blood type is negative for that type (Figure 14).
- For example, if a precipitate forms for A and Rh for a person’s blood sample, then they are A+. If a precipitate forms for A and B, but not Rh, that person’s blood type is AB- If the blood samples do not react to any serum, then that blood sample is O-.
- Take screenshots and record the data in the Suspect Information Sheet before moving on to the next suspect, as the results will not save once a new test begins.
- Repeat steps 2-5 for each suspect by clicking on the Next Test option at the bottom right of the page.
- The blood found at the scene of the crime was A+. Test Roy’s blood to make sure that the blood at the scene is the killer’s blood and not Roy’s blood.
- Use the collected data in the Suspect Information Sheet to determine if any suspect’s blood type correlates with the killer’s blood type.
3. Fingerprint Test
The procedure below was previously performed and videoed by TAs. Watch this video for this portion of the Crime Scene Investigation. Download the video if it does not play in the browser. The steps are outlined for your information only.
- Following a procedure that is similar to the one used in the Forensics Academy, carefully obtain the murder weapon (the shard of glass).
- Carefully apply the iron filament at the rim of the glass weapon and try not to waste filament.
- Using the magnetic wand, carefully remove the excess filament.
- Fingerprint residue should be left behind after removing the excess filament; if not, repeat steps 2 and 3.
- Observe the fingerprint, noting its features and type. Determine which person on the Suspect Information Sheet has that type. Figure 15 shows a clearer view of the fingerprint.
Determine who the killer is using all the information gathered in the above three analyses and the data provided in the Suspect Information Sheet. Tell a TA who you think the killer is before leaving the lab.
The lab work is now complete. Refer to the Assignment section for the instructions to prepare the lab report.
Assignment
Team Lab Report
Follow the lab report guidelines laid out in the EG1004 Writing Style Guide in the Technical Writing section of the manual. Use the outline below to write this report.
- Discuss the importance of biomedical forensics
- Explain the three steps of DNA extraction
- Describe the most common types of fingerprints
- Explain what causes different blood types
- Justify the use of salt, soap, and alcohol in the extraction procedure (i.e. discuss the important properties of DNA directly having an impact on the extraction procedure)
- Discuss how knowing the victim's blood type helped find the killer
- Clearly describe the procedural steps that were carried out in lab
- Include the steps carried out by the TA in the videos
- Include pictures of the extracted DNA, fingerprint on the murder weapon, and drug present in the victim’s blood
- Include the blood type of the killer
- How would the procedure change if meat was used instead of fruit in the DNA extraction process?
- Discuss improvements that could be made to the experiment
Remember: Lab notes must be taken. Experimental details are easily forgotten unless written down. EG1004 Lab Notes paper can be downloaded and printed from the EG1004 Website. Use the lab notes to write the Procedure section of the lab report. At the end of each lab, a TA will scan the lab notes and upload them to the Lab Documents section of the EG1004 Website. One point of extra credit is awarded if the lab notes are attached at the end of the lab report. Keeping careful notes is an essential component of all scientific practice.
Team PowerPoint Presentation
Follow the presentation guidelines laid out in the EG1004 Lab Presentation Format in the Technical Presentations section of the manual. When preparing the presentation, consider the following points.
- Rely heavily on graphics and pictures
- Make sure the experimental work is described simply and thoroughly
- Discuss the real-life application of DNA extraction
- Demonstrate clear understanding of each procedural step carried out and why it worked
References
Science Probeware & Experiment Software for Teachers. (2020, April 02). Retrieved July 29, 2020, from https://www.vernier.com/
Thermo Fisher Scientific - US. (n.d.). Retrieved July 29, 2020, from https://www.thermofisher.com/in/en/home.html
TRANSFORMING FOR TOMORROW WEBINARS. (2020, January 06). Retrieved July 29, 2020, from https://nasco.com/
SCQ. (2020, December 06). Retrieved July 29, 2020, from https://www.scq.ubc.ca/
Laboratory 3: Amplification of Your Mitochondrial DNA by PCR. (n.d.). Retrieved July 29, 2020, from http://hhmi.princeton.edu/lab-protocols/past-protocols/25-laboratory-3-amplification-of-your-mitochondrial-dna-by-pcr
http://porpax.bio.miami.edu/~cmallery/255/255chem/255chemistry.htm
http://library.thinkquest.org/20465/DNAstruct.html
Find and share research. (n.d.). Retrieved July 29, 2020, from https://www.researchgate.net/
| ||||||||